As I had stated in my last post, the discussion on the Make Me Taller boards seemed to have lead to me finding another rather revolutionary way to possibly increase a person’s height.
The original Discussion was started HERE but after a few extra clicks of linkes, I found myself on this discussion where one of the profiles started to talk about an idea that was very foreign to me, and seemed also very hopeful located HERE.
…..The spine is a very delicate and dynamic aspect of our body. The Call To Torsos is for a compilation of research that may help discover or advance methods for minimally-invasive or non-invasive methods for increasing Sitting Height via lengthening the Introvertebral Discs.
The major challenge is, to increase the spine length you would need a minimally-invasive or non-invasive method for increasing the height of the Vertebrae (Bone – not at all ideal) or of the Introvertebral Discs (Fibrocartilage – very ideal).
You do not want to increase the bone length because this is likely to be more difficult and there will be increased load on the Introvertebral Discs. The Introvertebral Discs provide critical cushioning and mobility between the Vertebrae and they degenerate with age. If you could increase the height of the Introvertebral Discs, you would be combating the degenerative effects of age which lead to spinal injury, you would have a healthier spine less prone to injury and you would increase height.
The optimistic aspect is 1: science has approached a period where it is trying to master growing Bone and Fibrocartilage among many other biological tissues and 2: 24 Vertebrae are not fused and thus 25 Discs have reasonable potential to be manipulated http://www.giantscientific.com/height_gain_exercise.html , which means that only a very small increase in height needs to be applied to each disc (e.g. 0.5cm or half of the width of your pinky nail Multiplied By 25 = 12.5cm = about 5 inches or 4.92 inches to be exact).
While some research is being applied to methods for manipulating the body in completely non-invasive ways such as systemic oral consumption or even approaches similar to magnetics or radio waves etc, my theory is that in order to specifically increase height of the discs in a practical manner which will not have negative side effects on the rest of the body one will likely need minimally-invasive injections directly to the disc.
I imagine a solution such as Ultrasound-Guided Injections of some sort of growth agent, probably best administered while inverting on a specially designed Inversion Table. This also needs to be done in a manner where the disc does actually grow the correct amount in the correct direction and in a uniform/proportional manner so that there is no bulging and interference with the nerves.
Nothing would likely be permanent, since the discs degenerate with age or the agent that may be found to do this may simply “inflate” the disc rather than actually grow it. However, there could be varying degrees of maintenance depending on application. It would be Controlled Introvertebral Fibrocartilage Hypertrophy and/or Hyperplasia or something technically close to that – not a permanent implant or anything incredibly invasive.
While this may seem too extreme at first though, the reality is Leg Lengthening is accepted here and it is a violent breaking of the largest bones in the body followed by lengthening with a lot of tearing/stretching of soft tissues, consolidating and soft tissue healing and rehab for months and years; whereas this is simply injections that could be done in a single day and actually improve the quality/safety/mobility/health of your spine and possibly provide about 5 inches in height. This could be similar to women getting botox injections to inflate their lips or actors/models getting botox to compensate for indented scars etc. You could even observe which discs would benefit best from an increase via the Ultrasound and make sure to take care of them first.
Me: On the GrownGrowth’s next post, he goes even further to explain the method/ technique to increase torso height
So, let’s talk about the anatomy and method again in order to address this perspective of yours.
Method:
Controlled Introvertebral Fibrocartilage Hypertrophy and/or Hyperplasia, which I theorize is to be done by Ultrasound-Guided Injections while inverted.
Methodological application on anatomy:
Introvertebral Fibrocartilage Hypertrophy and/or Hyperplasia means…
Controlled lengthening of the cartilage (type: Fibrocartilage), commonly referred to as Spinal Discs, in between the vertebrae (Introvertebral) by causing the Discs to grow in a controlled manner.
Essentially, we recognize that growth plates that help bones grow are also made of cartilage (type: Hyaline) before they become bone; and that the Spinal Discs could greatly benefit from reasonable addition in size for health and anti-degeneration while also increasing height (old people partly get shorter from Spinal Disc Degeneration, the Discs provide flexibility and shock absorbing …etc).
Real example of the direct dynamic effect of the method on anatomy:
24 Vertebrae are not fused and thus 25 Discs have reasonable potential to be manipulated http://www.giantscientific.com/height_gain_exercise.html , which means that only a very small increase in height needs to be applied to each disc (e.g. 0.5cm or half of the width of your pinky nail Multiplied By 25 = 12.5cm = about 5 inches or 4.92 inches to be exact).
*If you pay attention to the Sitting Height : Leg Height : Total Height Ratios, you’ll realize that 5 inches is going to be towards the rare extreme of maximum lengthening … much like only about 2 people on this forum have lengthened their legs 6 inches while most are thinking about 2-3 inches.
Real example put into perspective of the systemic or indirect effect on the anatomy:
With leg lengthening, there is usually 1 lateral cut in the bone per segment (rarely 2). This means that all of the lengthening is occurring at one “pressure point” or “load point” so to speak. So to compare LL of bones to the lengthening of the spine, like you were trying to do, this would be as if you only enlarged 1 of the 25 discs by 5 inches, whereas I am talking about enlarging each of the 25 discs a very small amount (again, even 5 inches is only 0.5cm lengthening per disc or the width of half of your pinky nail – think of how incredibly small that is for 5 inches of effect). While the overall increase is large, the “load dispersion” spread proportionally by only 0.5cm makes this very practical for 1: how much each disc can grow, 2: how much this effects surrounding anatomy because there is no large gap created in any one area rather the effect itself is so proportional that it is “almost systemic”.
(Common sense, plus see my skin experiment and rope analogy below!)
More importantly, is the very elaborate understanding of how each of those organs are secured to the body and/or secured to each other.
So, let’s try to put the effect on the organs in perspective with all of this:
First, we’ll eliminate what we should:
Quote
the anus, from which fecal wastes are excreted. Finally, the pelvic region houses both the male and female reproductive organs.
These organs are in the fused section of the spine, and thus the only load that is going to be applied here is approximately 0.5cm for the 1 and only disc directly connected at the top and a similarly small load at the point where the intestine connects etc. It will not be the collective load, but the load in the specific area of connection.
Quote
In the upper chest, the heart and lungs are protected by the rib cage
So, we know for sure that the rib cage is firmly attached to the spine. We need to be incredibly concerned that when lengthening the spine the heart and lungs are still going to be as protected as they should be. What I do not understand, nor know how to certainly find out without dissecting a cadaver or talking with an experienced surgeon, is exactly how the heart and lungs are secured and thus how they would move in relation to the spine and ribs moving.
What is good to know is, a healthy body is already quite flexible. For example, when somebody performs a reverse arch type of pose:
….the spine is effectively lengthening in some respect by the angle it bends even though there isn’t a height increase – maybe 1-3+ inches total – because of the flexibility of the Introvertebral Discs – and the ribs are moving greatly, yet the lungs and heart are also moving with the ribs and they are still protected even though there is a better angle for a spear to enter the heart.
Now, it is important to remember that even though a total height increase in my example of 5inches, the “load dispersion” means that the ribs themselves are not elevated by 5 inches! The bottom rib will elevate approximately 0.5cm and the ribs gaps will be stretched approximately 0.5cm per rib as they are connected to each vertebrae and they still have connective tissue in the gaps for protection!
Regardless, as long as the heart and lungs are attached well enough – which I think/hope they are – they will simply move with the ribs (not need to grow or stretch at all) and be protected the same.
Quote
the abdomen contains the majority of organs responsible for digestion: the stomach, where food is broken down; the pancreas and liver, which respectively produce enzymes and bile necessary for digestion; the kidneys, which filter wastes from blood before excretion as urine; the large and small intestines, which extract nutrients from food;
Again, I do not know how all of these are connected, but they are going to have a very small load of 0.5cm put on them very proportionately.
I’m quite sure this is not going to matter at all for the intestines. I bet it would make you a bit more regular because it would stretch them out a bit (don’t you notice how well your digest and dump after abdomen stretching?)! The small intestine is about 23 feet long and the large intestine is about 5 feet long (plenty of room the lengthen 0.5cm proportionally along their surface area against the spine)!
Likewise, the rest of the organs could very possibly handle the proportional load covered only for their surface area in relation to the spine.
So, what essentially remains is the skin and the muscles of the back and abdomen and their connective fascia/tissues. This, combined with gravity, should be the biggest resistance how I currently imagine. However, Hyperplasia and/or Hypertrophy can very logically potentially overcome this!
Now, to try to give an example of how easy the skin will surely adapt, do this:
Pinch the skin on your bicep between your thumb and index finger, grip it and pull it as far out from the arm as possible. You’ll probably get an inch of stretch easily. This is because the “load” is dispersed proportionally throughout the already-flexible skin.
However, let go. Now, think of only the small area of skin that your pinched for a grip and try to spread that small area of skin out the same inch or so that the skin collectively extended from you arm. You can’t because there is too much “load” on too small of an area, thus you have to tear the skin and cause it to grow new skin to stretch the same inch or so as the first experiment! However, you can stretch that small area about 0.5cm before the skin would have to tear, which helps make the point that each Introvertebral Disc could likely be manipulated by about 0.5cm!
In theory, the muscle and all connective tissue itself may adapt the same!
One other example by analogy is, imagine an old rope. Quality rope is usually made by 3 core strands interwoven, and each core strand has many micro strands of thread. An old rope loosens over time, with the core strands unweaving and the micro strands unweaving. The longer the rope is, the more the length difference. A very long and loosened rope, when pulled from each end, may increase in length by 5 inches; however, that is only because each micro strand and core strand is tightening proportionally throughout it’s entire length and no smaller segment of the rope could be lengthened 5 inches without cutting the rope.
I don’t think I am completely wrong. I may be completely correct. At worst I am partly correct. There certainly isn’t enough reason to be deterred based on this premise alone.
Quote from: Soho on October 01, 2008, 09:16:13 PM
And for me it’s clearly not with today technology that we will make it.
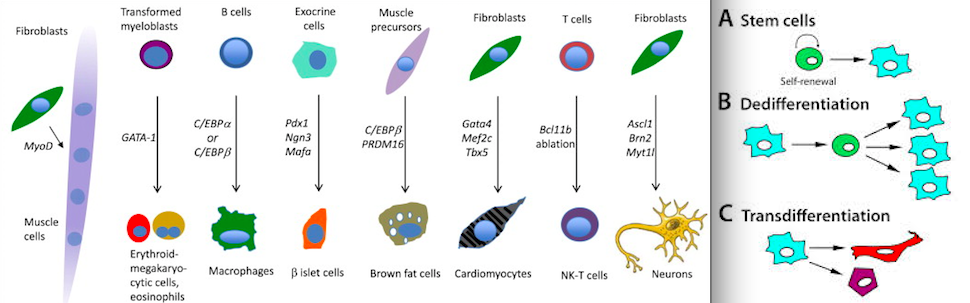
No, it is very possible with current technology. If we don’t already have all the pieces of the puzzle, we are very close. You still think stem cell progress is science fiction and you’re way off. HGH, IGF, MGF, BGH and Adult and Embryonic stem cells and more are all very well understood – though maybe not perfectly – and we are growing all sorts of tissues in vitro and directly on some animals and body builders are growing tissues on themselves etc.