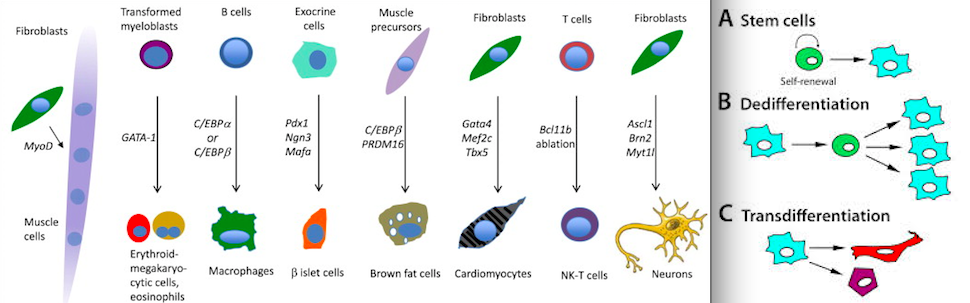
I wrote some about calcium secretions(which relate to ATP oscillations) here. TGF-Beta forms pre-chondrogenic mesenchymal condensation(which is what creates the growth plate) via ATP oscillations. ATP oscillations also play a role in FGF and Shh mesenchymal condensations.
Analysis of proteins showing differential changes during ATP oscillations in chondrogenesis.
“Prechondrogenic condensation is a critical step for skeletal pattern formation{ie form growth plates}. ATP oscillations play an essential role in prechondrogenic condensation because they induce oscillatory secretion. We examined how differential changes in proteins are implicated in ATP oscillations during chondrogenesis by using liquid chromatography/mass spectrometry. A number of proteins involved in ATP synthesis/consumption, catabolic/anabolic processes, actin dynamics, cell migration and adhesion were detected at either the peak or the trough of ATP oscillations, which implies that these proteins have oscillatory expression patterns that are coupled to ATP oscillations. On the basis of the results, we suggest that (1) the oscillatory expression of proteins involved in ATP synthesis/consumption and catabolic/anabolic processes can contribute to the generation or maintenance of ATP oscillations and that (2) the oscillatory expression of proteins involved in actin dynamics, cell migration and adhesion plays key roles in prechondrogenic condensation by inducing collective adhesion and migration in cooperation with ATP oscillations.”
So we can compare the proteins altered in ATP oscillations to those in LSJL to help see if LSJL induces similar ATP oscillations as those in growth plate chondrogenesis.
“ATP oscillations depend on Ca2+ dynamics.”
“We used the prechondrogenic ATDC5 cell line”<-It would be more ideal if they used normal mesenchymal stem cells as that’s what we’re trying to use to create new growth plates rather than the ATDC5 pre-chondrogenic cells that are like the pre-cursor cells in the Ring of LaCroix.
Peak versus Trough of ATP oscillations. Genes up and downregulated in LSJL are mentioned in {}
| No | Peak | Trough |
|---|---|---|
| 1 | Obg-like ATPase 1 | Suppression of tumorigenicity 5 protein |
| 2 | Proteasome subunit alpha type-7 | Glucosylceramidase |
| 3 | Serine/threonine-protein phosphatase PP1-beta catalytic subunit | Alpha-crystallin B chain |
| 4 | Hydroxymethylglutaryl-CoA lyase | Cytochrome c, somatic |
| 5 | Paired box protein Pax-7{down as Pax7a} | Arylsulfatase B |
| 6 | Arf-GAP with SH3 domain, ANK repeat and PH domain-containing protein 2 | Translationally controlled tumour protein |
| 7 | Lysosomal protective protein | Actin-related protein 2/3 complex subunit 4 |
| 8 | Protein canopy homolog 4 | StAR-related lipid transfer protein 3 |
| 9 | GRB2-associated-binding protein 3 | Transmembrane protein 5 |
| 10 | Platelet-activating factor acetylhydrolase IB subunit beta | Proteolipid protein 2 |
| 11 | Proteasome subunit alpha type-6 | Ribonuclease inhibitor |
| 12 | 60S ribosomal protein L30 | Glycyl-tRNA synthetase |
| 13 | 26S proteasome non-ATPase regulatory subunit 12 | Oxysterol-binding protein-related protein 3 |
| 14 | Protein CREG1 | Forkhead box protein N3 |
| 15 | Inorganic pyrophosphatase | Nuclear transport factor 2 |
| 16 | Myeloid cell nuclear differentiation antigen-like protein | LIM and SH3 domain protein 1 |
| 17 | Sorting nexin-3 | Serine/arginine-rich splicing factor 3 |
| 18 | Developmental pluripotency-associated protein 4 | SAP domain-containing ribonucleoprotein |
| 19 | BUD13 homolog | Growth/differentiation factor 7 |
| 20 | cAMP-responsive element-binding protein-like 2 | Prostaglandin E synthase 3 |
| 21 | Tau-tubulin kinase 2 | Ornithine aminotransferase |
| 22 | Thymosin beta-10 | Protein DGCR14 |
| 23 | 40S ribosomal protein S20 | Vang-like protein 2 |
| 24 | Copper transport protein ATOX1 | Mitochondrial import inner membrane translocase subunit Tim13 |
| 25 | Pulmonary surfactant-associated protein D | Zinc transporter ZIP8 |
| 26 | 40S ribosomal protein S11 | Thrombospondin-3 |
| 27 | 26S protease regulatory subunit 10B | Kinase suppressor of Ras 1 |
| 28 | Haematological and neurological expressed 1-like protein | Protein KIAA0284 |
| 29 | Beta-2-microglobulin | 40S ribosomal protein S4, X isoform |
| 30 | Vasohibin-2 | Galactokinase |
| 31 | Ubiquilin-1 | Scavenger mRNA-decapping enzyme DcpS |
| 32 | Nucleolysin TIAR | PHD and RING finger domain-containing protein 1 |
| 33 | Cleavage stimulation factor subunit 2 | Acyl-coenzyme A thioesterase 1 |
| 34 | Heparanase | Aspartyl-tRNA synthetase, cytoplasmic |
| 35 | Drebrin-like protein | TOM1-like protein 1 |
| 36 | Interferon regulatory factor 2-binding protein-like | Zinc finger CCHC domain-containing protein 14 |
| 37 | Golgin subfamily A member 2 | Anaphase-promoting complex subunit 5 |
| 38 | Potassium-transporting ATPase alpha chain 1 | Armadillo repeat-containing protein 8 |
| 39 | Collagen alpha-1(XVI) chain{up Col16a1} | Coiled-coil-helix-coiled-coil-helix domain-containing protein 2 |
| 40 | SWI/SNF complex subunit SMARCC1 | UPF0568 protein C14orf166 homolog |
| 41 | Cleavage stimulation factor subunit 3 | Putative potassium channel regulatory protein |
| 42 | Centrosomal protein of 170 kDa | MAM domain-containing glycosylphosphatidyl -nositol anchor protein 1 |
| 43 | Zinc finger protein 609 | UPF0160 protein MYG1 |
| 44 | CD44 antigen | Nucleoredoxin-like protein 1 |
| 45 | Transcription factor COE2 | Osteoclast-stimulating factor 1 |
| 46 | Heterogeneous nuclear ribonucleoproteins C1/C2 | Pyrroline-5-carboxylate reductase 1 |
| 47 | Inosine triphosphate pyrophosphatase | 40S ribosomal protein S5 |
| 48 | Latexin | Protein SET |
| 49 | DNA replication licencing factor MCM2 | Succinyl-CoA ligase [GDP-forming] subunit beta, mitochondrial |
| 50 | Motile sperm domain-containing protein 2 | Anoctamin-1 |
| 51 | Group XVI phospholipase A2 | Annexin A7 |
| 52 | 26S protease regulatory subunit 7 | Fibrillin-1 |
| 53 | Sodium- and chloride-dependent betaine transporter | GDP-mannose 4,6 dehydratase |
| 54 | Sorcin | Homogentisate 1,2-dioxygenase |
| 55 | Small ubiquitin-related modifier 2 | BTB/POZ domain-containing protein KCTD12 |
| 56 | Dihydropyrimidinase-related protein 3 | Mitogen-activated protein kinase kinase kinase MLK4 |
| 57 | ATP-dependent RNA helicase DDX39A | NEDD4 family-interacting protein 1 |
| 58 | Eukaryotic translation initiation factor 3 subunit L | Secretory carrier-associated membrane protein 1 |
| 59 | Far upstream element-binding protein 2 | Alpha-2,8-sialyltransferase 8E |
| 60 | Glypican-5 | Cytochrome b-c1 complex subunit Rieske, mitochondrial |
| 61 | Intraflagellar transport protein 74 homolog | Serine/threonine-protein phosphatase 6 regulatory subunit 2 |
| 62 | Uncharacterized protein KIAA0141 | Interleukin-15 receptor subunit alpha |
| 63 | Paralemmin-3 | Cell cycle progression protein 1 |
| 64 | Peroxiredoxin-6 | Uncharacterized protein C4orf36 homolog |
| 65 | Proteasome subunit alpha type-2 | Leukocyte surface antigen CD47 |
| 66 | Proteasome subunit beta type-3 | Cyclic nucleotide-gated cation channel beta-3 |
| 67 | Reticulocalbin-2 | Fibroblast growth factor 14 |
| 68 | TAR DNA-binding protein 43 | Hypoxanthine-guanine phosphoribosyltransferase |
| 69 | Ubiquitin-conjugating enzyme E2 K | Interferon alpha-12 |
| 70 | ATP-binding cassette subfamily D member 4 | Leucine-rich repeat and IQ domain-containing protein 3 |
| 71 | Atlastin-3 | Neuroplastin |
| 72 | Voltage-dependent l-type calcium channel subunit beta-4 | Rab proteins geranylgeranyltransferase component A 1 |
| 73 | Coronin-6 | SHC-transforming protein 4 |
| 74 | Endonuclease/exonuclease/phosphatase family domain-containing protein 1 | Vacuolar protein sorting-associated protein 29 |
| 75 | Exocyst complex component 1 | Plakophilin-3 |
| 76 | Insulin-like growth factor-binding protein 5 | Transmembrane protein 223 |
| 77 | Lysosomal alpha-mannosidase | Clathrin light chain A |
| 78 | Nucleus accumbens-associated protein 1 | Apoptosis-inducing factor 1, mitochondrial |
| 79 | Phosphoprotein associated with glycosphingolipid-enriched microdomains 1 | Protein-S-isoprenylcysteine O-methyltransferase |
| 80 | 26S proteasome non-ATPase regulatory subunit 7 | Aspartyl aminopeptidase |
| 81 | SH3 domain-binding glutamic acid-rich-like protein 3 | Tumour protein p53-inducible nuclear protein 1 |
| 82 | Sepiapterin reductase | Nuclear pore complex protein Nup54 |
| 83 | Tripeptidyl-peptidase 1 | ETS domain-containing protein Elk-4 |
| 84 | V-type proton ATPase subunit d 1 | Sideroflexin-3 |
| 85 | V-type proton ATPase subunit G 1 | Ferric-chelate reductase 1 |
| 86 | Xaa-Pro aminopeptidase 1 | Williams–Beuren syndrome chromosomal region 14 protein homolog |
| 87 | 4F2 cell-surface antigen heavy chain | ER degradation-enhancing alpha-mannosidase-like 1 |
| 88 | Disrupted in schizophrenia 1 homolog | Pumilio domain-containing protein KIAA0020 |
| 89 | Biglycan | 25-Hydroxyvitamin d-1 alpha hydroxylase, mitochondrial |
| 90 | Apolipoprotein A-I-binding protein | Uncharacterized protein C8orf42 homolog |
| 91 | Biliverdin reductase A | Solute carrier family 12 member 5 |
| 92 | RNA/RNP complex-1-interacting phosphatase | Inosine-5′-monophosphate dehydrogenase 2 |
| 93 | Homeobox protein Hox-A5 | BRCA1-associated RING domain protein 1 |
| 94 | Dynein light chain 1, cytoplasmic | Growth hormone-regulated TBC protein 1 |
| 95 | Beta-1,3-galactosyl-O-glycosyl-glycoprotein beta-1,6-N-acetylglucosaminyltransferase | Calcitonin gene-related peptide 2 |
| 96 | Arf-GAP with dual PH domain-containing protein 2 | ATP-dependent RNA helicase DDX1 |
| 97 | ATP-binding cassette subfamily A member 8-B | Trans-2,3-enoyl-CoA reductase |
| 98 | RING finger protein unkempt-like | Laminin subunit gamma-2 |
| 99 | Ankyrin-2 | |
| 100 | Complement C1q subcomponent subunit C | |
| 101 | Probable ATP-dependent RNA helicase DDX59 | |
| 102 | Homeobox protein Nkx-2.2 | |
| 103 | Heterogeneous nuclear ribonucleoprotein M | |
| 104 | FtsJ methyltransferase domain-containing protein 1 | |
| 105 | Uncharacterized protein C1orf141 homolog | |
| 106 | Cell division protein kinase 5 | |
| 107 | Protein tyrosine phosphatase type IVA 1 | |
| 108 | Uncharacterized protein C4orf34 homolog | |
| 109 | Ephrin-A5 | |
| 110 | Cytochrome b-c1 complex subunit 7 | |
| 111 | Endoplasmic reticulum lectin 1 | |
| 112 | Bifunctional apoptosis regulator | |
| 113 | Coiled-coil domain-containing protein 27 | |
| 114 | Long-chain specific acyl-CoA dehydrogenase, mitochondrial | |
| 115 | Iroquois-class homeodomain protein IRX-5 | |
| 116 | Endoplasmic reticulum mannosyl- oligosaccharide 1,2-alpha-mannosidase | |
| 117 | Cysteine protease ATG4B | |
| 118 | Galactoside 2-alpha-l-fucosyltransferase 3 | |
| 119 | Apolipoprotein A-II | |
| 120 | Kelch-like protein 24 | |
| 121 | Serine/threonine-protein kinase Nek5 | |
| 122 | Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma isoform |
With categories
| Function | Peak | Trough |
|---|---|---|
| ATP synthesis | V-type proton ATPase subunit d 1 | Cytochrome c |
| V-type proton ATPase subunit G 1 | Cytochrome b-c1 complex subunit Rieske | |
| Potassium-transporting ATPase alpha chain 1 | ||
| Cytochrome b-c1 complex subunit 7 | ||
| Phosphorylation | Tau-tubulin kinase 2 | Galactokinase |
| Cell division protein kinase 5 | Mitogen-activated protein kinase kinase kinase MLK4 | |
| Serine/threonine-protein kinase Nek5 | ||
| Phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit gamma isoform | ||
| Dephosphorylation | Serine/threonine-protein phosphatase PP1-beta catalytic subunit | Serine/threonine-protein phosphatase 6 regulatory subunit 2 |
| Inorganic pyrophosphatase, | ||
| Inosine triphosphate pyrophosphatase | ||
| Protein tyrosine phosphatase type IVA 1 | ||
| RNA/RNP complex-1-interacting phosphatase | ||
| Biosynthetic process | StAR-related lipid transfer protein 3 | |
| Prostaglandin E synthase 3 | ||
| Pyrroline-5-carboxylate reductase 1 | ||
| Inosine-5′-monophosphate dehydrogenase 2 | ||
| Trans-2,3-enoyl-CoA reductase | ||
| Catabolic process | Proteasome subunit alpha type-7 | Ferric-chelate reductase 1 |
| Proteasome subunit alpha type-6 | ER degradation-enhancing alpha-mannosidase-like 1 | |
| 26S protease regulatory subunit 10B | 25-Hydroxyvitamin d-1 alpha hydroxylase | |
| 26S protease regulatory subunit 7 | ||
| Proteasome subunit alpha type-2 | ||
| Proteasome subunit beta type-3 | ||
| Endoplasmic reticulum lectin 1 | ||
| Long-chain specific acyl-CoA dehydrogenase | ||
| Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase | ||
| Biliverdin reductase A | ||
| Tripeptidyl-peptidase 1 | ||
| Xaa-Pro aminopeptidase 1 | ||
| Heparanase | ||
| Actin dynamics | Thymosin beta-10 | Actin-related protein 2/3 complex subunit 4 |
| Alpha-crystallin B chain (stabilize) | ||
| LIM and SH3 domain protein 1 (stabilize) | ||
| Cell adhesion | Collagen alpha-1(XVI) chain | Laminin subunit gamma-2 |
| Plakophilin-3 | ||
| Neuroplastin, leukocyte surface antigen CD47 | ||
| Thrombospondin-3 | ||
| Cell migration | Platelet-activating factor acetylhydrolase IB subunit beta | MAM domain-containing glycosylphosphatidylinositol anchor protein 1 |
| CD44 antigen | ||
| Homeobox protein Hox-A5 | ||
| Cell division protein kinase 5 | ||
| Ephrin-A5 |
Comparison to LSJL genes to be done.
” it was found that many proteins involved in cell adhesion, such as laminin subunit gamma-2, plakophilin-3, neuroplastin, thrombospondin-3 and leukocyte surface antigen CD47, were detected only at the troughs of ATP oscillations whereas many proteins involved in cell migration, such as platelet-activating factor acetylhydrolase IB subunit beta, CD44 antigen, homeobox protein Hox-A5, cell division protein kinase 5, and ephrin-A5, were detected only at the peaks of ATP oscillations”
” cell–cell adhesion and cell movement are stimulated at the troughs and peaks of ATP oscillations, respectively, leading to synchronized oscillations of cellular migration during chondrogenesis”
” A nucleation and growth mechanism in which a critical-size aggregate (nuclei) is required for subsequent growth can be applicable to the cellular aggregation process. Therefore, we propose that prechondrogenic condensation proceeds by a nucleation-growth mechanism. This could explain the reason why the oscillatory expression patterns of proteins that are involved in actin dynamics, cell migration and adhesion are required for prechondrogenic condensation. Thermodynamically, a barrier exists in free energy; this barrier occurs at a certain critical size at the initial stage of condensation. Thus, aggregates smaller than the critical size are unstable owing to energy loss due to surface tension, whereas aggregates larger than the critical size grow irreversibly owing to the energy gain due to cellular adhesions surpassing the energy loss, thereby ultimately forming large cellular aggregates. Therefore, the oscillatory expression patterns of proteins involved in actin dynamics, cell migration and adhesion result in collective migration and adhesion, which aggregates cells collectively within a limited time and then drives more efficient formation of the nuclei than random migration and adhesion. Therefore, coordinated oscillatory expression of the proteins is crucial during the initial step of prechondrogenic condensation.”
“platelets are aggregate into clusters at the site of an injury to the skin or blood vessels.”<-Note that blood is involved in distraction osteogenesis